 Nonlinear extensions of the LASSO exist such as modeling a binary outcome. LASSO is similar to OLS with constraints and produces a stable and interpretable model. LASSO is a combination of ridge regression and subset selection developed to improve OLS by shrinking the coefficient values and setting some equal to zero. In the case of a large number of predictors, OLS has difficulty selecting the subset of predictors that appears to be the most important or to have the strongest effects. Ordinary least squares (OLS) is known to estimate coefficients with small bias but inflated variance. Two important components of variable selection are prediction accuracy and interpretation. In an effort to improve variable selection, Tibshirani developed the least absolute shrinkage and selection operator (LASSO), a penalized likelihood approach, for linear regression. The research objective of this study is to develop prediction tools primarily methods for variable selection and dimension reduction in a GWAS. The tendency for analyzing genotype data is to use GLM and univariate tests however, these models perform poorly when analyzing high-dimensional data. Genetic data is used to find genetic variants that are associated with rheumatoid arthritis risk (or other diseases) through the use of statistical modeling. There is a demand for statistical techniques to handle large volumes of data, particularly in the area of genetics. The highly dense genetic marker data from the rheumatoid arthritis study and the published reports about the study provide an ideal empirical dataset for developing and testing extensions of dimension-reduction methods. We can apply such techniques to determine whether multiple marker pathways and gene-gene interactions are associated with the disease of interest. Dimension-reduction techniques are a powerful tool because they provide a summary measure of massive amounts of data. Recently, the focus has shifted to GWAS, where the emphasis can be placed on assessing whether multiple markers function together rather than depending on univariate tests and generalized linear models (GLM). have paved new directions for dimension-reduction techniques and broadened the area to other applications of prediction, including genetics.įor this paper, we explore extensions of currently existing dimension-reduction methods and variable-selection methods related to genome-wide association studies (GWAS) single-nucleotide polymorphism (SNP) selection and gene-gene interactions for application to the disease classification problem based on genetic data. 
For example, during the last decade, Li, Tibshirani, and Efron et al. This has called for broadening of the area of research in dimension-reduction techniques to provide methods for prediction and variable selection. Technical advances have enabled the collection of massive high-dimensional datasets in such studies. There is a demand for statistical techniques capable of handling large volumes of data in genetic studies. The goal of this paper is to develop and evaluate prediction methods and tools for genome-wide association studies, particularly for variable selection and dimension reduction. Based on our results, the PCA-LASSO method shows promise in identifying gene-gene interactions, and, at this time we suggest using it with other conventional approaches, such as generalized linear models, to narrow down genetic signals. 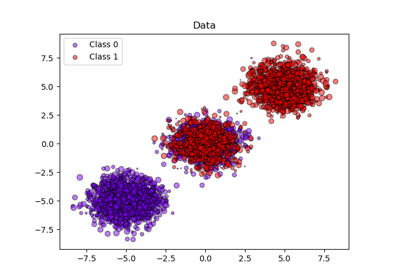
We demonstrated these methods with the Genetic Analysis Workshop 16 rheumatoid arthritis genome-wide association study data and our results identified a few gene-gene signals. This method was compared to placing the raw SNP values into the LASSO and the logistic model with individual gene-gene interaction. We have extended the PCA-LASSO approach using the bootstrap to estimate the standard errors and confidence intervals of the LASSO coefficient estimates. 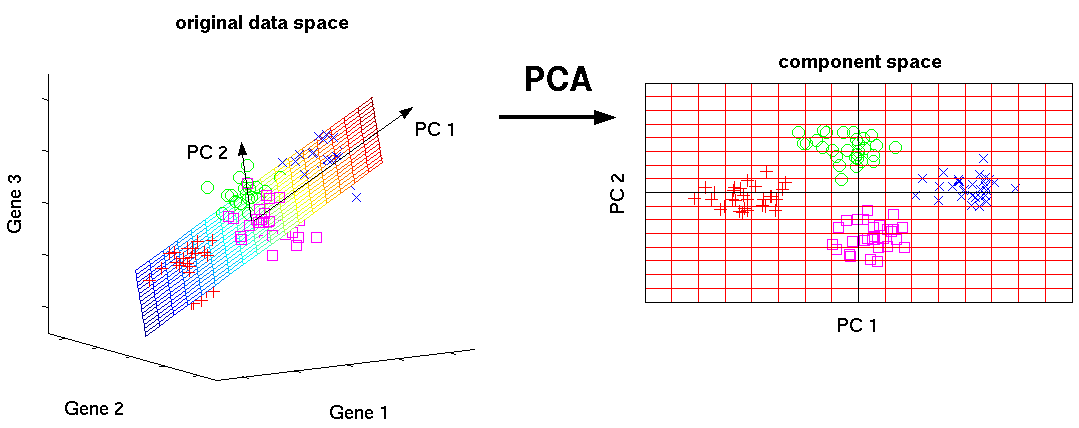
The interaction of the gene PCA scores were placed into LASSO to determine whether any gene-gene signals exist. A PCA was used to first reduce the dimension of the single-nucleotide polymorphisms (SNPs) within each gene. We propose an approach that uses principal-component analysis (PCA) and least absolute shrinkage and selection operator (LASSO) to identify gene-gene interaction in genome-wide association studies. Variable selection in genome-wide association studies can be a daunting task and statistically challenging because there are more variables than subjects. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
